Abstract
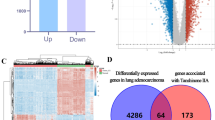
Extracellular regulated protein kinases 1/2 (ERK1/2) are key members of multiple signaling pathways, including the ErbB axis. Ectopic ERK1/2 activation contributes to various types of cancer, especially drug resistance to inhibitors of RTK, RAF and MEK, and specific ERK1/2 inhibitors are scarce. In this study, we identified a potential novel covalent ERK inhibitor, Laxiflorin B, which is a herbal compound with anticancer activity. However, Laxiflorin B is present at low levels in herbs; therefore, we adopted a semi-synthetic process for the efficient production of Laxiflorin B to improve the yield. Laxiflorin B induced mitochondria-mediated apoptosis via BAD activation in non-small-cell lung cancer (NSCLC) cells, especially in EGFR mutant subtypes. Transcriptomic analysis suggested that Laxiflorin B inhibits amphiregulin (AREG) and epiregulin (EREG) expression through ERK inhibition, and suppressed the activation of their receptors, ErbBs, via a positive feedback loop. Moreover, mass spectrometry analysis combined with computer simulation revealed that Laxiflorin B binds covalently to Cys-183 in the ATP-binding pocket of ERK1 via the D-ring, and Cys-178 of ERK1 through non-inhibitory binding of the A-ring. In a NSCLC tumor xenograft model in nude mice, Laxiflorin B also exhibited strong tumor suppressive effects with low toxicity and AREG and EREG were identified as biomarkers of Laxiflorin B efficacy. Finally, Laxiflorin B-4, a C-6 analog of Laxiflorin B, exhibited higher binding affinity for ERK1/2 and stronger tumor suppression. These findings provide a new approach to tumor inhibition using natural anticancer compounds.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Lavoie H, Gagnon J, Therrien M. ERK signalling: a master regulator of cell behaviour, life and fate. Nat Rev Mol Cell Biol. 2020;21:607–32.
Samatar AA, Poulikakos PI. Targeting RAS-ERK signalling in cancer: promises and challenges. Nat Rev Drug Discov. 2014;13:928–42.
Arcila ME, Drilon A, Sylvester BE, Lovly CM, Borsu L, Reva B, et al. MAP2K1 (MEK1) mutations define a distinct subset of lung adenocarcinoma associated with smoking. Clin Cancer Res. 2015;21:1935–43.
Degirmenci U, Wang M, Hu J. Targeting aberrant RAS/RAF/MEK/ERK signaling for cancer therapy. Cells. 2020;9:198.
Brenan L, Andreev A, Cohen O, Pantel S, Kamburov A, Cacchiarelli D, et al. Phenotypic characterization of a comprehensive set of MAPK1/ERK2 missense mutants. Cell Rep. 2016;17:1171–83.
Roskoski R Jr. Targeting ERK1/2 protein-serine/threonine kinases in human cancers. Pharmacol Res. 2019;142:151–68.
Germann UA, Furey BF, Markland W, Hoover RR, Aronov AM, Roix JJ, et al. Targeting the MAPK signaling pathway in cancer: promising preclinical activity with the novel selective ERK1/2 inhibitor BVD-523 (Ulixertinib). Mol Cancer Ther. 2017;16:2351–63.
Sullivan RJ, Infante JR, Janku F, Wong DJL, Sosman JA, Keedy V, et al. First-in-class ERK1/2 inhibitor ulixertinib (BVD-523) in patients with MAPK mutant advanced solid tumors: results of a phase I dose-escalation and expansion study. Cancer Discov. 2018;8:184–95.
Blake JF, Burkard M, Chan J, Chen H, Chou KJ, Diaz D, et al. Discovery of (S)-1-(1-(4-Chloro-3-fluorophenyl)-2-hydroxyethyl)-4-(2-((1-methyl-1H-pyrazol-5-y l)amino)pyrimidin-4-yl)pyridin-2(1H)-one (GDC-0994), an extracellular signal-regulated kinase 1/2 (ERK1/2) inhibitor in early clinical development. J Med Chem. 2016;59:5650–60.
Sunaga N, Kaira K, Imai H, Shimizu K, Nakano T, Shames DS, et al. Oncogenic KRAS-induced epiregulin overexpression contributes to aggressive phenotype and is a promising therapeutic target in non-small-cell lung cancer. Oncogene. 2013;32:4034–42.
Bhagwat SV, McMillen WT, Cai S, Zhao B, Whitesell M, Shen W, et al. ERK inhibitor LY3214996 targets ERK pathway-driven cancers: a therapeutic approach toward precision medicine. Mol Cancer Ther. 2020;19:325–36.
Varga A, Soria JC, Hollebecque A, LoRusso P, Bendell J, Huang SA, et al. A first-in-human phase I study to evaluate the ERK1/2 inhibitor GDC-0994 in patients with advanced solid tumors. Clin Cancer Res. 2020;26:1229–36.
Kirouac DC, Schaefer G, Chan J, Merchant M, Orr C, Huang SA, et al. Clinical responses to ERK inhibition in BRAF(V600E)-mutant colorectal cancer predicted using a computational model. NPJ Syst Biol Appl. 2017;3:14.
Moschos SJ, Sullivan RJ, Hwu WJ, Ramanathan RK, Adjei AA, Fong PC, et al. Development of MK-8353, an orally administered ERK1/2 inhibitor, in patients with advanced solid tumors. JCI Insight. 2018;3:e92352.
Morris EJ, Jha S, Restaino CR, Dayananth P, Zhu H, Cooper A, et al. Discovery of a novel ERK inhibitor with activity in models of acquired resistance to BRAF and MEK inhibitors. Cancer Discov. 2013;3:742–50.
Liu M, Wang WG, Sun HD, Pu JX. Diterpenoids from Isodon species: an update. Nat Prod Rep. 2017;34:1090–140.
Wang WG, Du X, Li XN, Wu HY, Liu X, Shang SZ, et al. New bicyclo[3.1.0]hexane unit ent-kaurane diterpene and its seco-derivative from Isodon eriocalyx var. laxiflora. Org Lett. 2012;14:302–5.
Sun HD, Lin ZW, Niu FD, Shen PQ, Pan LT, Lin LZ, et al. Diterpenoids from Isodon eriocalyx var. laxiflora. Phytochemistry. 1995;38:1451–5.
van Zundert GCP, Rodrigues JPGLM, Trellet M, Schmitz C, Kastritis PL, Karaca E, et al. The HADDOCK2.2 web server: user-friendly integrative modeling of biomolecular complexes. J Mol Biol. 2016;428:720–5.
Rao S, Gurbani D, Du G, Everley RA, Browne CM, Chaikuad A, et al. Leveraging compound promiscuity to identify targetable cysteines within the kinome. Cell Chem Biol. 2019;26:818–29.e9.
Case DA, Ben-Shalom IY, Brozell SR, Cerutti DS, Cheatham TEI, Cruzeiro VWD, et al. AMBER 2018, San Francisco: University of California; 2018.
Maier JA, Martinez C, Kasavajhala K, Wickstrom L, Hauser KE, Simmerling C. FF14SB: improving the accuracy of protein side chain and backbone parameters from FF99SB. J Chem Theory Comput. 2015;11:3696–713.
Wang J, Wolf RM, Caldwell JW, Kollman PA, Case DA. Development and testing of a general amber force field. J Comput Chem. 2004;25:1157–74.
Jorgensen WL, Chandrasekhar J, Madura JD, Impey RW, Klein ML. Comparison of simple potential functions for simulating liquid water. J Chem Phys. 1983;79:926–35.
Qiu D, Shenkin PS, Hollinger FP, Still WC. The GB/SA continuum model for solvation. a fast analytical method for the calculation of approximate born radii. J Phys Chem A. 1997;101:3005–14.
Aronchik I, Dai Y, Labenski M, Barnes C, Jones T, Qiao L, et al. Efficacy of a covalent ERK1/2 inhibitor, CC-90003, in KRAS-mutant cancer models reveals novel mechanisms of response and resistance. Mol Cancer Res. 2019;17:642–54.
Ohori M, Kinoshita T, Yoshimura S, Warizaya M, Nakajima H, Miyake H. Role of a cysteine residue in the active site of ERK and the MAPKK family. Biochem Biophys Res Commun. 2007;353:633–7.
Niu XM, Li SH, Li ML, Zhao QS, Mei SX, Na Z, et al. Cytotoxic ent-kaurane diterpenoids from Isodon eriocalyx var. laxiflora. Planta Med. 2002;68:528–33.
Elangovan IM, Vaz M, Tamatam CR, Potteti HR, Reddy NM, Reddy SP. FOSL1 promotes Kras-induced lung cancer through amphiregulin and cell survival gene regulation. Am J Respir Cell Mol Biol. 2018;58:625–35.
Lechtenberg BC, Mace PD, Sessions EH, Williamson R, Stalder R, Wallez Y, et al. Structure-guided strategy for the development of potent bivalent ERK inhibitors. ACS Med Chem Lett. 2017;8:726–31.
Garai Á, Zeke A, Gógl G, Törő I, Fördős F, Blankenburg H, et al. Specificity of linear motifs that bind to a common mitogen-activated protein kinase docking groove. Sci Signal 2012;5:ra74.
Chatterjee P, Botello-Smith WM, Zhang H, Qian L, Alsamarah A, Kent D, et al. Can relative binding free energy predict selectivity of reversible covalent inhibitors? J Am Chem Soc. 2017;139:17945–52.
Ward RA, Colclough N, Challinor M, Debreczeni JE, Eckersley K, Fairley G, et al. Structure-guided design of highly selective and potent covalent inhibitors of ERK1/2. J Med Chem. 2015;58:4790–801.
Ohori M, Kinoshita T, Okubo M, Sato K, Yamazaki A, Arakawa H, et al. Identification of a selective ERK inhibitor and structural determination of the inhibitor-ERK2 complex. Biochem Biophys Res Commun. 2005;336:357–63.
Zhao S, Qiu ZX, Zhang L, Li WM. Prognostic values of ERK1/2 and p-ERK1/2 expressions for poor survival in non-small cell lung cancer. Tumour Biol. 2015;36:4143–50.
Blake JF, Gaudino JJ, De Meese J, Mohr P, Chicarelli M, Tian H, et al. Discovery of 5,6,7,8-tetrahydropyrido[3,4-d]pyrimidine inhibitors of Erk2. Bioorg Med Chem Lett. 2014;24:2635–9.
Chiang CY, Pan CC, Chang HY, Lai MD, Tzai TS, Tsai YS, et al. SH3BGRL3 protein as a potential prognostic biomarker for urothelial carcinoma: a novel binding partner of epidermal growth factor receptor. Clin Cancer Res. 2015;21:5601–11.
Deng Y, Shipps GW Jr, Cooper A, English JM, Annis DA, Carr D, et al. Discovery of novel, dual mechanism ERK inhibitors by affinity selection screening of an inactive kinase. J Med Chem. 2014;57:8817–26.
Leproult E, Barluenga S, Moras D, Wurtz JM, Winssinger N. Cysteine mapping in conformationally distinct kinase nucleotide binding sites: application to the design of selective covalent inhibitors. J Med Chem. 2011;54:1347–55.
Kohler J, Zhao Y, Li J, Gokhale PC, Tiv HL, Knott AR, et al. ERK inhibitor LY3214996-based treatment strategies for RAS-driven lung cancer. Mol Cancer Ther. 2021;20:641–54.
Plotnikov A, Flores K, Maik-Rachline G, Zehorai E, Kapri-Pardes E, Berti DA, et al. The nuclear translocation of ERK1/2 as an anticancer target. Nat Commun. 2015;6:6685.
Hu S, Xie Z, Onishi A, Yu X, Jiang L, Lin J, et al. Profiling the human protein-DNA interactome reveals ERK2 as a transcriptional repressor of interferon signaling. Cell. 2009;139:610–22.
Tee WW, Shen SS, Oksuz O, Narendra V, Reinberg D. Erk1/2 activity promotes chromatin features and RNAPII phosphorylation at developmental promoters in mouse ESCs. Cell. 2014;156:678–90.
Madak-Erdogan Z, Lupien M, Stossi F, Brown M, Katzenellenbogen BS. Genomic collaboration of estrogen receptor alpha and extracellular signal-regulated kinase 2 in regulating gene and proliferation programs. Mol Cell Biol. 2011;31:226–36.
Li Y, Cheng HS, Chng WJ, Tergaonkar V. Activation of mutant TERT promoter by RAS-ERK signaling is a key step in malignant progression of BRAF-mutant human melanomas. Proc Natl Acad Sci USA. 2016;113:14402–7.
Lee MH, Yanagawa J, Tran L, Walser TC, Bisht B, Fung E, et al. FRA1 contributes to MEK-ERK pathway-dependent PD-L1 upregulation by KRAS mutation in premalignant human bronchial epithelial cells. Am J Transl Res. 2020;12:409–27.
Cho MC, Choi HS, Lee S, Kim BY, Jung M, Park SN, et al. Epiregulin expression by Ets-1 and ERK signaling pathway in Ki-ras-transformed cells. Biochem Biophys Res Commun. 2008;377:832–7.
Sunaga N, Kaira K. Epiregulin as a therapeutic target in non-small-cell lung cancer. Lung Cancer. 2015;6:91–8.
Singh B, Coffey RJ. Trafficking of epidermal growth factor receptor ligands in polarized epithelial cells. Annu Rev Physiol. 2014;76:275–300.
Yang Y, Zhang Z, Li S, Ye X, Li X, He K. Synergy effects of herb extracts: pharmacokinetics and pharmacodynamic basis. Fitoterapia. 2014;92:133–47.
Acknowledgements
The authors would like to thank Dr. Jessica Tamanini of ETediting for editing the manuscript prior to submission and the Instrumental Analysis Center of Shenzhen University for the support. This work was supported by the Natural Science Foundation of Guangdong Province (No. 2021A1515011046, 2021A1515010996), the Guangdong Provincial Science and Technology Program (No. 2017B030301016), the Regional Joint Fund of Guangdong Province (Grant No. 2019B1515120080), the Shenzhen Municipal Government of China (No. JCYJ20210324093408024), the Shenzhen Key Medical Discipline Construction Fund (No. SZXK060).
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: DZ, LZZ and CYC. Chemical compound synthesis: JRH and MY. Conducted most of the experiments and analyzed the data: MZ, CYC, JRH, CLC, DMP, TTZ, HY and TX. Construction of plasmids and lentivirus packaging: MZ, CYC. and CLC. Animal studies: CYC and FY. Performed the LC–MS/MS and proteomic analysis: QQH, ZW and YDZ. Performed the computer simulation of Laxiflorin B and ERK1/2 crystal structures: JZ. Planning and discussion of the project: DZ and LZZ. Supervised the entire project: YCC, DZ and LZZ. Wrote the manuscript, designed the layout of figures and tables: CYC, DZ, ZGL, and JZ.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chiang, CY., Zhang, M., Huang, J. et al. A novel selective ERK1/2 inhibitor, Laxiflorin B, targets EGFR mutation subtypes in non-small-cell lung cancer. Acta Pharmacol Sin 45, 422–435 (2024). https://doi.org/10.1038/s41401-023-01164-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41401-023-01164-w